注意
转到末尾以下载完整示例代码或通过 JupyterLite 或 Binder 在浏览器中运行此示例。
用类别似然比衡量分类性能#
此示例演示了 class_likelihood_ratios 函数,该函数计算正负似然比(LR+、LR-)以评估二元分类器的预测能力。正如我们将看到的,这些指标独立于测试集中的类别比例,这使得当用于研究的可用数据与目标应用具有不同的类别比例时,它们非常有用。
一个典型的应用是医学中的病例对照研究,其中类别几乎平衡,而一般人群中的类别失衡较大。在这种应用中,个体患目标疾病的先验概率可以选择为流行率,即在特定人群中发现受某种疾病影响的比例。后验概率则代表在测试结果为阳性的情况下,疾病确实存在的概率。
在本例中,我们首先讨论由类别似然比给出的先验和后验赔率之间的联系。然后,我们在一些受控场景中评估它们的行为。在最后一节中,我们将它们绘制为正类别流行率的函数。
# Authors: The scikit-learn developers
# SPDX-License-Identifier: BSD-3-Clause
先验 vs. 后验分析#
假设我们有一个人群,具有生理测量值 X,这些测量值有望作为疾病的间接生物标志物,以及实际疾病指标 y(地面真实值)。人群中的大多数人没有携带疾病,但少数人(在本例中约为10%)携带疾病。
from sklearn.datasets import make_classification
X, y = make_classification(n_samples=10_000, weights=[0.9, 0.1], random_state=0)
print(f"Percentage of people carrying the disease: {100 * y.mean():.2f}%")
Percentage of people carrying the disease: 10.37%
建立一个机器学习模型来诊断具有给定生理测量值的人是否可能携带目标疾病。为了评估模型,我们需要在保留的测试集上评估其性能。
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
然后我们可以拟合我们的诊断模型并计算正似然比,以评估该分类器作为疾病诊断工具的有用性。
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import class_likelihood_ratios
estimator = LogisticRegression().fit(X_train, y_train)
y_pred = estimator.predict(X_test)
pos_LR, neg_LR = class_likelihood_ratios(y_test, y_pred, replace_undefined_by=1.0)
print(f"LR+: {pos_LR:.3f}")
LR+: 12.617
由于正类别似然比远大于1.0,这意味着基于机器学习的诊断工具是有用的:在测试结果为阳性的情况下,疾病确实存在的后验赔率比先验赔率大12倍以上。
似然比的交叉验证#
我们评估在某些特定情况下类别似然比测量的可变性。
import pandas as pd
def scoring(estimator, X, y):
y_pred = estimator.predict(X)
pos_lr, neg_lr = class_likelihood_ratios(y, y_pred, replace_undefined_by=1.0)
return {"positive_likelihood_ratio": pos_lr, "negative_likelihood_ratio": neg_lr}
def extract_score(cv_results):
lr = pd.DataFrame(
{
"positive": cv_results["test_positive_likelihood_ratio"],
"negative": cv_results["test_negative_likelihood_ratio"],
}
)
return lr.aggregate(["mean", "std"])
我们首先验证上一节中使用的具有默认超参数的 LogisticRegression 模型。
from sklearn.model_selection import cross_validate
estimator = LogisticRegression()
extract_score(cross_validate(estimator, X, y, scoring=scoring, cv=10))
我们确认该模型是有用的:后验赔率比先验赔率大12到20倍。
相反,让我们考虑一个虚拟模型,它将输出随机预测,其赔率与训练集中的平均疾病流行率相似。
from sklearn.dummy import DummyClassifier
estimator = DummyClassifier(strategy="stratified", random_state=1234)
extract_score(cross_validate(estimator, X, y, scoring=scoring, cv=10))
在这里,两个类别似然比都与1.0兼容,这使得该分类器作为改进疾病检测的诊断工具毫无用处。
虚拟模型的另一个选择是始终预测最常见的类别,在这种情况下是“无疾病”。
estimator = DummyClassifier(strategy="most_frequent")
extract_score(cross_validate(estimator, X, y, scoring=scoring, cv=10))
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
No samples were predicted for the positive class and `positive_likelihood_ratio` is set to `np.nan`. Use the `replace_undefined_by` param to
缺乏阳性预测意味着没有真阳性也没有假阳性,导致 LR+ 未定义,绝不应解释为无限 LR+(分类器完美识别阳性病例)。在这种情况下,class_likelihood_ratios 函数返回 nan 并默认发出警告。事实上,LR- 的值帮助我们否决了这个模型。
当交叉验证高度不平衡且样本量少的数据时,可能会出现类似的情况:某些折叠中没有患病样本,因此在用于测试时不会输出真阳性或假阴性。在数学上,这会导致无限 LR+,也不应将其解释为模型完美识别阳性病例。这种情况会导致估计似然比的方差更高,但仍可解释为患病后验赔率的增加。
estimator = LogisticRegression()
X, y = make_classification(n_samples=300, weights=[0.9, 0.1], random_state=0)
extract_score(cross_validate(estimator, X, y, scoring=scoring, cv=10))
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
`positive_likelihood_ratio` is ill-defined and set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
`positive_likelihood_ratio` is ill-defined and set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
`positive_likelihood_ratio` is ill-defined and set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
`positive_likelihood_ratio` is ill-defined and set to `np.nan`. Use the `replace_undefined_by` param to
/home/circleci/project/sklearn/utils/_param_validation.py:218: UndefinedMetricWarning:
`positive_likelihood_ratio` is ill-defined and set to `np.nan`. Use the `replace_undefined_by` param to
关于流行率的不变性#
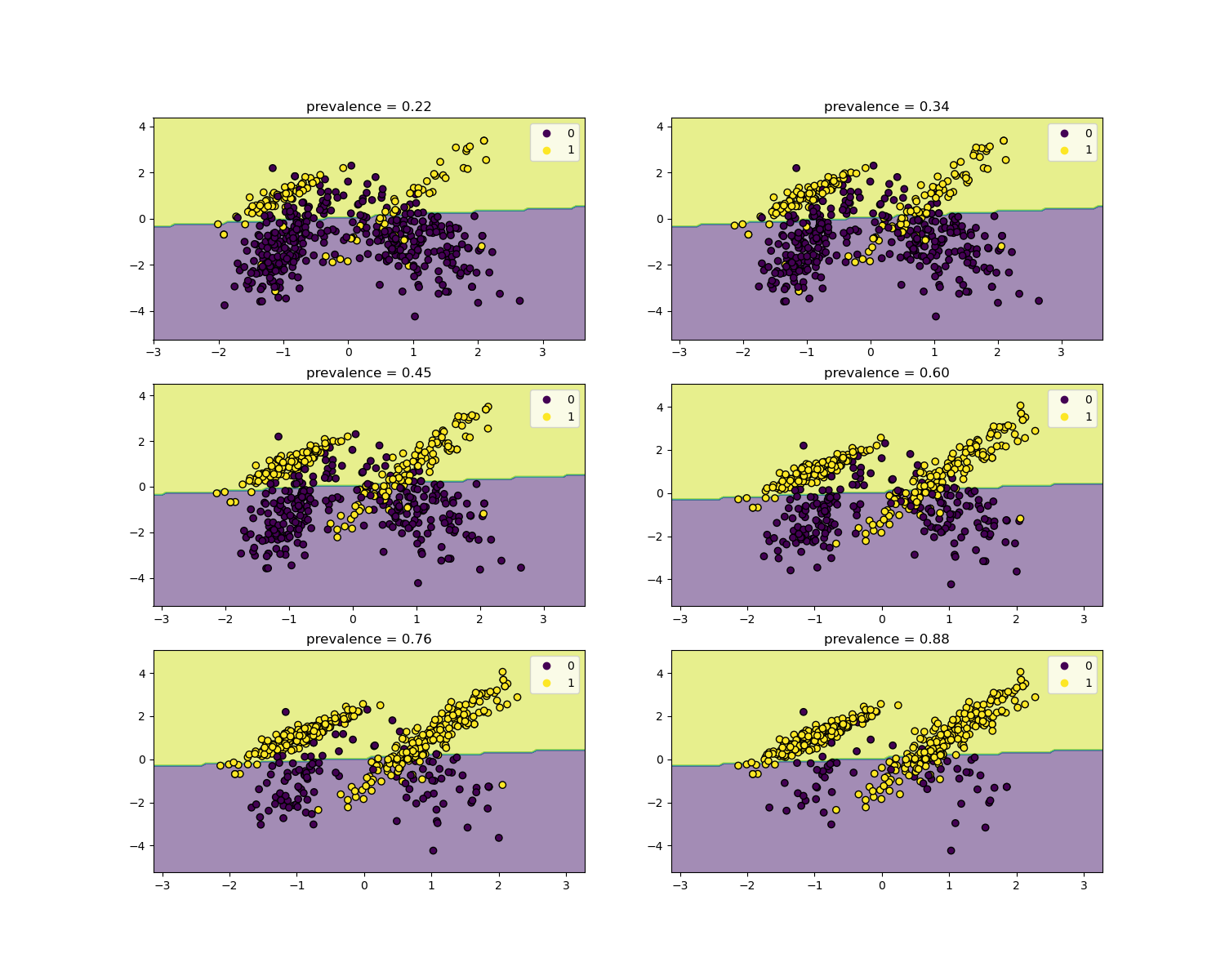
似然比独立于疾病流行率,可以在不同人群之间进行推断,无论是否存在类别失衡,只要对所有人群应用相同的模型。请注意,在下面的图中,决策边界是恒定的(有关不平衡类别的决策边界研究,请参阅 SVM:不平衡类别的分离超平面)。
在这里,我们在流行率为50%的病例对照研究上训练一个 LogisticRegression 基础模型。然后在一系列不同流行率的人群上进行评估。我们使用 make_classification 函数以确保数据生成过程始终与下面图中所示相同。标签 1 对应于正类别“疾病”,而标签 0 代表“无疾病”。
from collections import defaultdict
import matplotlib.pyplot as plt
import numpy as np
from sklearn.inspection import DecisionBoundaryDisplay
populations = defaultdict(list)
common_params = {
"n_samples": 10_000,
"n_features": 2,
"n_informative": 2,
"n_redundant": 0,
"random_state": 0,
}
weights = np.linspace(0.1, 0.8, 6)
weights = weights[::-1]
# fit and evaluate base model on balanced classes
X, y = make_classification(**common_params, weights=[0.5, 0.5])
estimator = LogisticRegression().fit(X, y)
lr_base = extract_score(cross_validate(estimator, X, y, scoring=scoring, cv=10))
pos_lr_base, pos_lr_base_std = lr_base["positive"].values
neg_lr_base, neg_lr_base_std = lr_base["negative"].values
我们现在将展示每个流行率级别的决策边界。请注意,我们只绘制了原始数据的一个子集,以便更好地评估线性模型的决策边界。
fig, axs = plt.subplots(nrows=3, ncols=2, figsize=(15, 12))
for ax, (n, weight) in zip(axs.ravel(), enumerate(weights)):
X, y = make_classification(
**common_params,
weights=[weight, 1 - weight],
)
prevalence = y.mean()
populations["prevalence"].append(prevalence)
populations["X"].append(X)
populations["y"].append(y)
# down-sample for plotting
rng = np.random.RandomState(1)
plot_indices = rng.choice(np.arange(X.shape[0]), size=500, replace=True)
X_plot, y_plot = X[plot_indices], y[plot_indices]
# plot fixed decision boundary of base model with varying prevalence
disp = DecisionBoundaryDisplay.from_estimator(
estimator,
X_plot,
response_method="predict",
alpha=0.5,
ax=ax,
)
scatter = disp.ax_.scatter(X_plot[:, 0], X_plot[:, 1], c=y_plot, edgecolor="k")
disp.ax_.set_title(f"prevalence = {y_plot.mean():.2f}")
disp.ax_.legend(*scatter.legend_elements())
我们定义一个用于引导的函数。
def scoring_on_bootstrap(estimator, X, y, rng, n_bootstrap=100):
results_for_prevalence = defaultdict(list)
for _ in range(n_bootstrap):
bootstrap_indices = rng.choice(
np.arange(X.shape[0]), size=X.shape[0], replace=True
)
for key, value in scoring(
estimator, X[bootstrap_indices], y[bootstrap_indices]
).items():
results_for_prevalence[key].append(value)
return pd.DataFrame(results_for_prevalence)
我们使用引导对每个流行率的基础模型进行评分。
results = defaultdict(list)
n_bootstrap = 100
rng = np.random.default_rng(seed=0)
for prevalence, X, y in zip(
populations["prevalence"], populations["X"], populations["y"]
):
results_for_prevalence = scoring_on_bootstrap(
estimator, X, y, rng, n_bootstrap=n_bootstrap
)
results["prevalence"].append(prevalence)
results["metrics"].append(
results_for_prevalence.aggregate(["mean", "std"]).unstack()
)
results = pd.DataFrame(results["metrics"], index=results["prevalence"])
results.index.name = "prevalence"
results
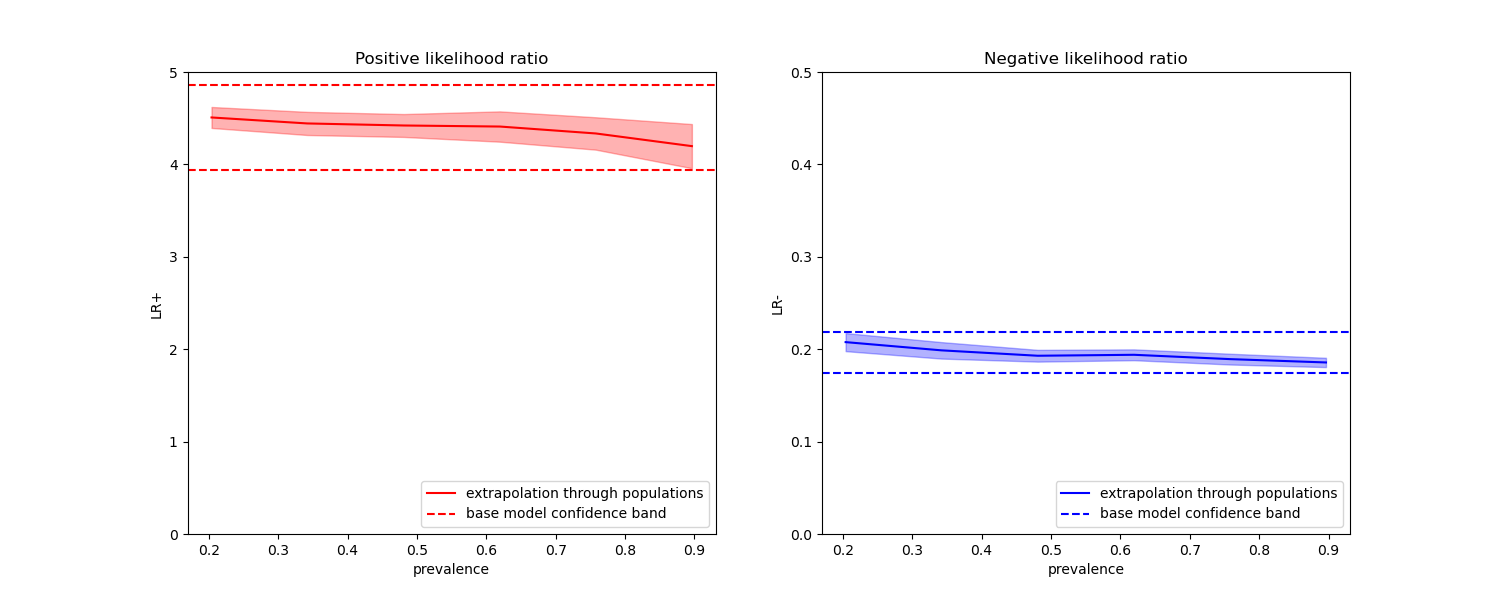
在下面的图中,我们观察到使用不同流行率重新计算的类别似然比在与平衡类别计算的似然比的一个标准差内确实是恒定的。
fig, (ax1, ax2) = plt.subplots(nrows=1, ncols=2, figsize=(15, 6))
results["positive_likelihood_ratio"]["mean"].plot(
ax=ax1, color="r", label="extrapolation through populations"
)
ax1.axhline(y=pos_lr_base + pos_lr_base_std, color="r", linestyle="--")
ax1.axhline(
y=pos_lr_base - pos_lr_base_std,
color="r",
linestyle="--",
label="base model confidence band",
)
ax1.fill_between(
results.index,
results["positive_likelihood_ratio"]["mean"]
- results["positive_likelihood_ratio"]["std"],
results["positive_likelihood_ratio"]["mean"]
+ results["positive_likelihood_ratio"]["std"],
color="r",
alpha=0.3,
)
ax1.set(
title="Positive likelihood ratio",
ylabel="LR+",
ylim=[0, 5],
)
ax1.legend(loc="lower right")
ax2 = results["negative_likelihood_ratio"]["mean"].plot(
ax=ax2, color="b", label="extrapolation through populations"
)
ax2.axhline(y=neg_lr_base + neg_lr_base_std, color="b", linestyle="--")
ax2.axhline(
y=neg_lr_base - neg_lr_base_std,
color="b",
linestyle="--",
label="base model confidence band",
)
ax2.fill_between(
results.index,
results["negative_likelihood_ratio"]["mean"]
- results["negative_likelihood_ratio"]["std"],
results["negative_likelihood_ratio"]["mean"]
+ results["negative_likelihood_ratio"]["std"],
color="b",
alpha=0.3,
)
ax2.set(
title="Negative likelihood ratio",
ylabel="LR-",
ylim=[0, 0.5],
)
ax2.legend(loc="lower right")
plt.show()
脚本总运行时间: (0 minutes 1.750 seconds)
相关示例