注意
点击此处下载完整的示例代码,或通过JupyterLite或Binder在浏览器中运行此示例。
t-SNE:不同困惑度值对形状的影响#
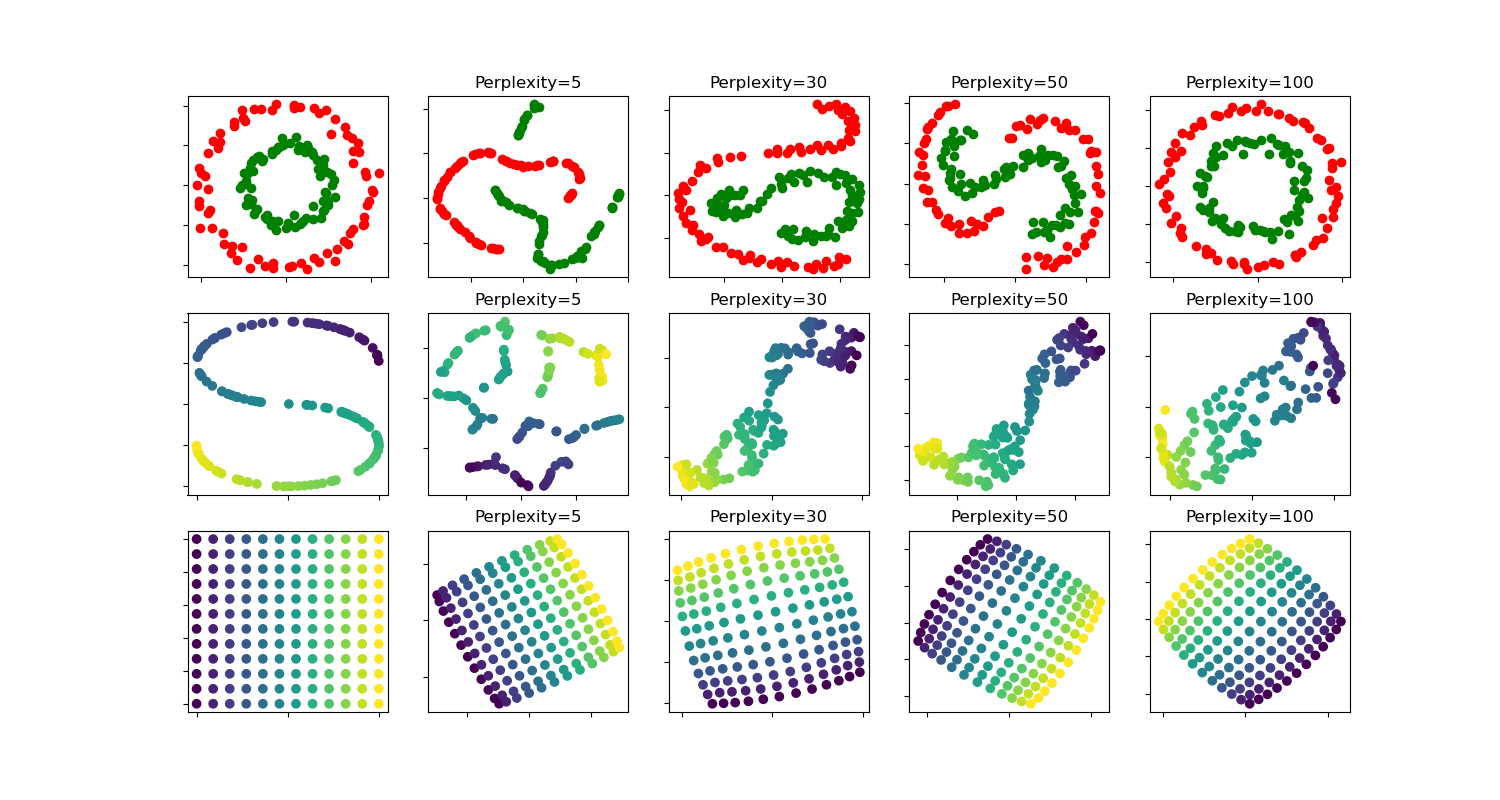
在两个同心圆和S曲线数据集上,针对不同的困惑度值,对t-SNE进行了说明。
我们观察到,随着困惑度值的增加,形状变得更加清晰。
簇的大小、距离和形状可能会根据初始化、困惑度值而变化,并不总是具有意义。
如下所示,对于较高的困惑度值,t-SNE能够找到两个同心圆的有意义的拓扑结构,但是圆的大小和距离与原始数据略有不同。与两个圆形数据集相反,即使对于较大的困惑度值,S曲线数据集上的形状在视觉上也与S曲线拓扑结构有所偏离。
更多详情,请参阅“如何有效使用t-SNE” https://distill.pub/2016/misread-tsne/,该文章很好地讨论了各种参数的影响,并提供了交互式图表来探索这些影响。
circles, perplexity=5 in 0.17 sec
circles, perplexity=30 in 0.28 sec
circles, perplexity=50 in 0.23 sec
circles, perplexity=100 in 0.26 sec
S-curve, perplexity=5 in 0.13 sec
S-curve, perplexity=30 in 0.2 sec
S-curve, perplexity=50 in 0.24 sec
S-curve, perplexity=100 in 0.23 sec
uniform grid, perplexity=5 in 0.2 sec
uniform grid, perplexity=30 in 0.27 sec
uniform grid, perplexity=50 in 0.27 sec
uniform grid, perplexity=100 in 0.27 sec
# Authors: The scikit-learn developers
# SPDX-License-Identifier: BSD-3-Clause
from time import time
import matplotlib.pyplot as plt
import numpy as np
from matplotlib.ticker import NullFormatter
from sklearn import datasets, manifold
n_samples = 150
n_components = 2
(fig, subplots) = plt.subplots(3, 5, figsize=(15, 8))
perplexities = [5, 30, 50, 100]
X, y = datasets.make_circles(
n_samples=n_samples, factor=0.5, noise=0.05, random_state=0
)
red = y == 0
green = y == 1
ax = subplots[0][0]
ax.scatter(X[red, 0], X[red, 1], c="r")
ax.scatter(X[green, 0], X[green, 1], c="g")
ax.xaxis.set_major_formatter(NullFormatter())
ax.yaxis.set_major_formatter(NullFormatter())
plt.axis("tight")
for i, perplexity in enumerate(perplexities):
ax = subplots[0][i + 1]
t0 = time()
tsne = manifold.TSNE(
n_components=n_components,
init="random",
random_state=0,
perplexity=perplexity,
max_iter=300,
)
Y = tsne.fit_transform(X)
t1 = time()
print("circles, perplexity=%d in %.2g sec" % (perplexity, t1 - t0))
ax.set_title("Perplexity=%d" % perplexity)
ax.scatter(Y[red, 0], Y[red, 1], c="r")
ax.scatter(Y[green, 0], Y[green, 1], c="g")
ax.xaxis.set_major_formatter(NullFormatter())
ax.yaxis.set_major_formatter(NullFormatter())
ax.axis("tight")
# Another example using s-curve
X, color = datasets.make_s_curve(n_samples, random_state=0)
ax = subplots[1][0]
ax.scatter(X[:, 0], X[:, 2], c=color)
ax.xaxis.set_major_formatter(NullFormatter())
ax.yaxis.set_major_formatter(NullFormatter())
for i, perplexity in enumerate(perplexities):
ax = subplots[1][i + 1]
t0 = time()
tsne = manifold.TSNE(
n_components=n_components,
init="random",
random_state=0,
perplexity=perplexity,
learning_rate="auto",
max_iter=300,
)
Y = tsne.fit_transform(X)
t1 = time()
print("S-curve, perplexity=%d in %.2g sec" % (perplexity, t1 - t0))
ax.set_title("Perplexity=%d" % perplexity)
ax.scatter(Y[:, 0], Y[:, 1], c=color)
ax.xaxis.set_major_formatter(NullFormatter())
ax.yaxis.set_major_formatter(NullFormatter())
ax.axis("tight")
# Another example using a 2D uniform grid
x = np.linspace(0, 1, int(np.sqrt(n_samples)))
xx, yy = np.meshgrid(x, x)
X = np.hstack(
[
xx.ravel().reshape(-1, 1),
yy.ravel().reshape(-1, 1),
]
)
color = xx.ravel()
ax = subplots[2][0]
ax.scatter(X[:, 0], X[:, 1], c=color)
ax.xaxis.set_major_formatter(NullFormatter())
ax.yaxis.set_major_formatter(NullFormatter())
for i, perplexity in enumerate(perplexities):
ax = subplots[2][i + 1]
t0 = time()
tsne = manifold.TSNE(
n_components=n_components,
init="random",
random_state=0,
perplexity=perplexity,
max_iter=400,
)
Y = tsne.fit_transform(X)
t1 = time()
print("uniform grid, perplexity=%d in %.2g sec" % (perplexity, t1 - t0))
ax.set_title("Perplexity=%d" % perplexity)
ax.scatter(Y[:, 0], Y[:, 1], c=color)
ax.xaxis.set_major_formatter(NullFormatter())
ax.yaxis.set_major_formatter(NullFormatter())
ax.axis("tight")
plt.show()
脚本总运行时间:(0分钟3.339秒)
相关示例